pypercolate HPC performance metrics¶
In this Section, we compute basic performance metrics of the percolate.hpc module.
Namely, we time the execution and measure memory consumption of graph creation with spanning_2d_grid(), of computing the convolution coefficients with _binomial_pmf(), and of performing a single run with bond_microcanonical_statistics().
Preamble¶
# configure plotting
%config InlineBackend.rc = {'figure.dpi': 300, 'savefig.dpi': 300, \
'figure.figsize': (6, 3), 'font.size': 12, \
'figure.facecolor': (1, 1, 1, 0)}
%matplotlib inline
import timeit
import matplotlib.pyplot as plt
import memory_profiler
import numpy as np
import percolate
plt.style.use('ggplot')
System sizes¶
We determine performance metrics for the following system sizes.
dimensions = 2 ** np.arange(3, 11)
np.save('dimensions.npy', dimensions)
print(dimensions)
[ 8 16 32 64 128 256 512 1024]
System information¶
This Section details the hardware, the operating system, and the Python environment that collects the performance metrics.
# $ pip install py-cpuinfo
from cpuinfo import cpuinfo
{
key: value
for key, value in cpuinfo.get_cpu_info().items()
if key in ['arch', 'brand', 'count', 'hz_advertised', 'l2_cache_size', 'vendor_id']
}
{'arch': 'X86_64',
'brand': 'Intel(R) Core(TM) i7-2620M CPU @ 2.70GHz',
'count': 4,
'hz_advertised': '2.7000 GHz',
'l2_cache_size': '4096 KB',
'vendor_id': 'GenuineIntel'}
import psutil
print("Total memory: {:.1f} GiB".format(psutil.virtual_memory().total / 1024 ** 3))
Total memory: 7.7 GiB
import os
import platform
import sys
import pip
import pprint
print(platform.platform())
'Linux-3.13.0-24-generic-x86_64-with-LinuxMint-17-qiana'
print(
platform.python_version(),
platform.python_implementation(),
platform.python_compiler(),
)
3.4.0 CPython GCC 4.8.2
pprint.pprint({
package.key: package.version
for package in pip.get_installed_distributions()
if package.key in [
'cython', 'future', 'ipython', 'matplotlib', 'memory-profiler',
'networkx', 'numpy', 'percolate', 'pip', 'scipy', 'simoa',
]
})
{'cython': '0.22.1',
'future': '0.15.0',
'ipython': '3.2.1',
'matplotlib': '1.4.3',
'memory-profiler': '0.33',
'networkx': '1.10',
'numpy': '1.9.2',
'percolate': '0.3.0.post0.dev10+gce92429.dirty',
'pip': '7.1.0',
'scipy': '0.16.0',
'simoa': '0.4.1'}
np.show_config()
openblas_info:
library_dirs = ['/usr/lib']
language = f77
libraries = ['openblas']
blas_mkl_info:
NOT AVAILABLE
mkl_info:
NOT AVAILABLE
openblas_lapack_info:
NOT AVAILABLE
lapack_opt_info:
include_dirs = ['/usr/include/atlas']
library_dirs = ['/usr/lib/atlas-base/atlas', '/usr/lib/atlas-base']
language = f77
libraries = ['lapack', 'f77blas', 'cblas', 'atlas']
define_macros = [('ATLAS_INFO', '"\"3.10.1\""')]
lapack_mkl_info:
NOT AVAILABLE
atlas_3_10_threads_info:
NOT AVAILABLE
blas_opt_info:
library_dirs = ['/usr/lib']
language = f77
libraries = ['openblas']
atlas_info:
include_dirs = ['/usr/include/atlas']
library_dirs = ['/usr/lib/atlas-base/atlas', '/usr/lib/atlas-base']
language = f77
libraries = ['lapack', 'f77blas', 'cblas', 'atlas']
define_macros = [('ATLAS_INFO', '"\"3.10.1\""')]
atlas_threads_info:
NOT AVAILABLE
atlas_3_10_info:
NOT AVAILABLE
Scripts¶
For each performance metric, we define and run a script in an independent process. Each script reads the system sizes from disk and writes the performance metrics back to disk. We use independent scripts and processes here as especially memory usage measurements seem to be volatile if executed in the same process, IPython session, or notebook.
%%python
import memory_profiler
import numpy as np
import percolate
def get_graph_memory(dimension):
mem_before = memory_profiler._get_memory(-1)
graph = percolate.spanning_2d_grid(dimension)
mem_after = memory_profiler._get_memory(-1)
return mem_after - mem_before
dimensions = np.load('dimensions.npy')
graph_mem = np.fromiter(
(
get_graph_memory(dimension)
for dimension in dimensions
),
dtype=np.float,
)
np.save('graph_mem.npy', graph_mem)
%%python
import memory_profiler
import numpy as np
import percolate
def get_convolution_memory(dimension):
mem_before = memory_profiler._get_memory(-1)
convolution_factors = percolate.percolate._binomial_pmf(
n=2 * dimension * (dimension - 1), p=0.5,
)
mem_after = memory_profiler._get_memory(-1)
return mem_after - mem_before
dimensions = np.load('dimensions.npy')
convolution_mem = np.fromiter(
(
get_convolution_memory(dimension)
for dimension in dimensions
),
dtype=np.float,
)
np.save('convolution_mem.npy', convolution_mem)
%%python
import timeit
import numpy as np
import percolate
dimensions = np.load('dimensions.npy')
graph_times = np.fromiter(
(
timeit.timeit(
stmt='percolate.spanning_2d_grid({})'.format(dimension),
setup='import percolate',
number=1,
)
for dimension in dimensions
),
dtype=np.float,
)
np.save('graph_times.npy', graph_times)
%%python
import timeit
import numpy as np
import percolate
dimensions = np.load('dimensions.npy')
convolution_stmt = """\
[
percolate.percolate._binomial_pmf(n={}, p=p)
for p in np.linspace(0.4, 0.6, num=1)
]
"""
convolution_times = np.fromiter(
(
timeit.timeit(
stmt=convolution_stmt.format(2 * dimension * (dimension - 1)),
setup='import percolate; import numpy as np',
number=1,
)
for dimension in dimensions
),
dtype=np.float,
)
np.save('convolution_times.npy', convolution_times)
%%python
import timeit
import numpy as np
dimensions = np.load('dimensions.npy')
run_stmt = """\
percolate.hpc.bond_microcanonical_statistics(
seed=42, **perc_graph
)
"""
run_setup = """\
import percolate
import percolate.hpc
perc_graph = percolate.percolate.percolation_graph(
graph=percolate.spanning_2d_grid({}),
spanning_cluster=True,
)
"""
run_times = np.fromiter(
(
timeit.timeit(
stmt=run_stmt,
setup=run_setup.format(dimension),
number=1,
)
for dimension in dimensions
),
dtype=np.float,
)
np.save('run_times.npy', run_times)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:302: RuntimeWarning: invalid value encountered in power
ret['moments'] += ret['max_cluster_size'] ** np.arange(5)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:286: RuntimeWarning: invalid value encountered in power
ret['moments'] -= weights[i] ** np.arange(5)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:286: RuntimeWarning: invalid value encountered in subtract
ret['moments'] -= weights[i] ** np.arange(5)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:297: RuntimeWarning: invalid value encountered in power
ret['moments'] += weight ** np.arange(5)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:297: RuntimeWarning: invalid value encountered in add
ret['moments'] += weight ** np.arange(5)
%%python
import memory_profiler
import numpy as np
import percolate
import percolate.hpc
def get_run_memory(dimension):
perc_graph = percolate.percolate.percolation_graph(
graph=percolate.spanning_2d_grid(dimension),
spanning_cluster=True,
)
mem_before = memory_profiler._get_memory(-1)
return (
max(memory_profiler.memory_usage(
(
percolate.hpc.bond_microcanonical_statistics,
[],
dict(seed=42, **perc_graph)
),
interval=.01,
)) - mem_before
)
dimensions = np.load('dimensions.npy')
run_mem = np.fromiter(
(
get_run_memory(dimension)
for dimension in dimensions
),
dtype=np.float,
)
np.save('run_mem.npy', run_mem)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:302: RuntimeWarning: invalid value encountered in power
ret['moments'] += ret['max_cluster_size'] ** np.arange(5)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:286: RuntimeWarning: invalid value encountered in power
ret['moments'] -= weights[i] ** np.arange(5)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:286: RuntimeWarning: invalid value encountered in subtract
ret['moments'] -= weights[i] ** np.arange(5)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:297: RuntimeWarning: invalid value encountered in power
ret['moments'] += weight ** np.arange(5)
/home/sorge/repos/sci/pypercolate/percolate/hpc.py:297: RuntimeWarning: invalid value encountered in add
ret['moments'] += weight ** np.arange(5)
graph_mem = np.load('graph_mem.npy')
convolution_mem = np.load('convolution_mem.npy')
graph_times = np.load('graph_times.npy')
convolution_times = np.load('convolution_times.npy')
run_times = np.load('run_times.npy')
run_mem = np.load('run_mem.npy')
Performance plots¶
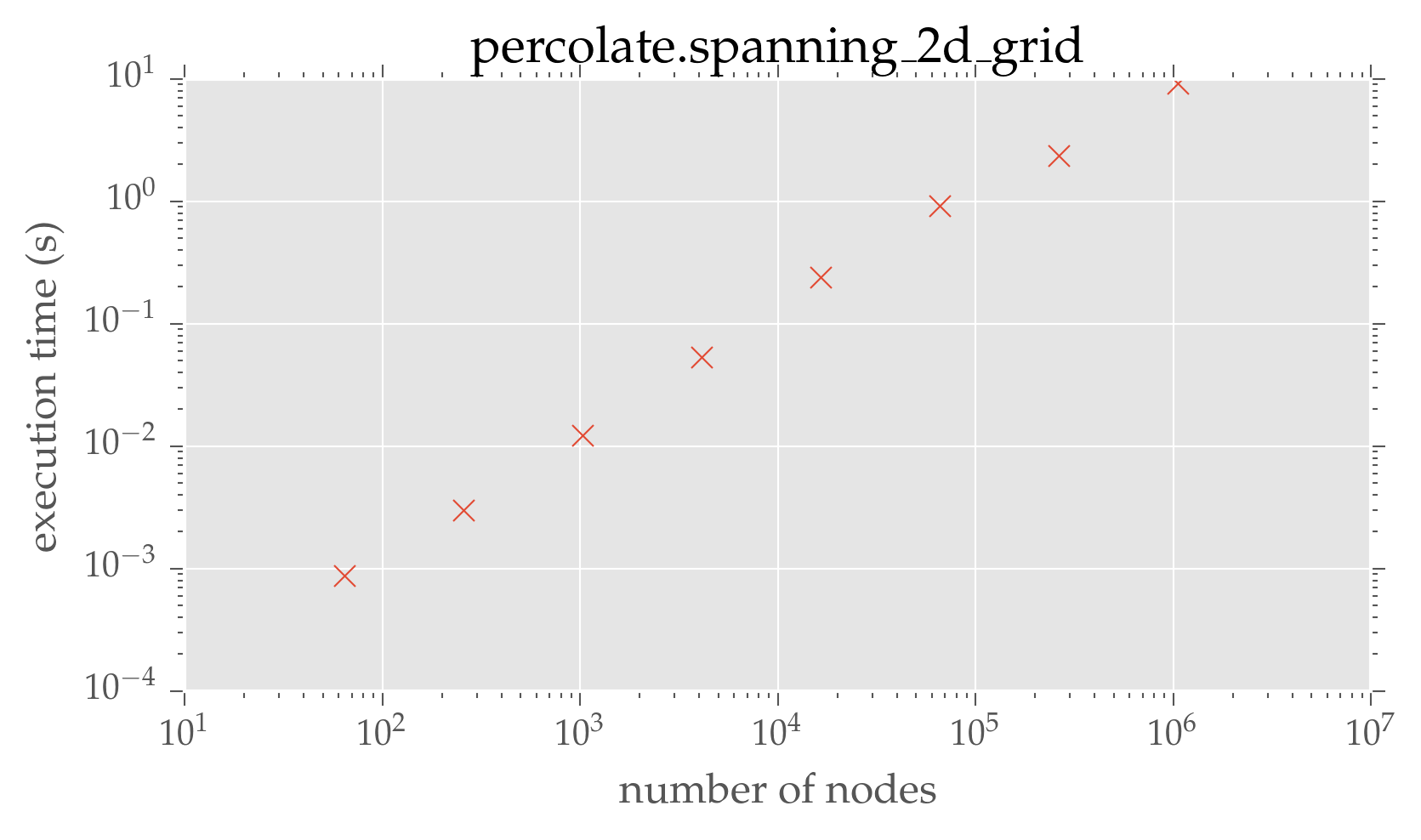
plt.loglog(dimensions ** 2, graph_times, 'x')
plt.xlabel(r'number of nodes')
plt.ylabel(r'execution time (s)')
plt.title(r'percolate.spanning\_2d\_grid')
plt.show()
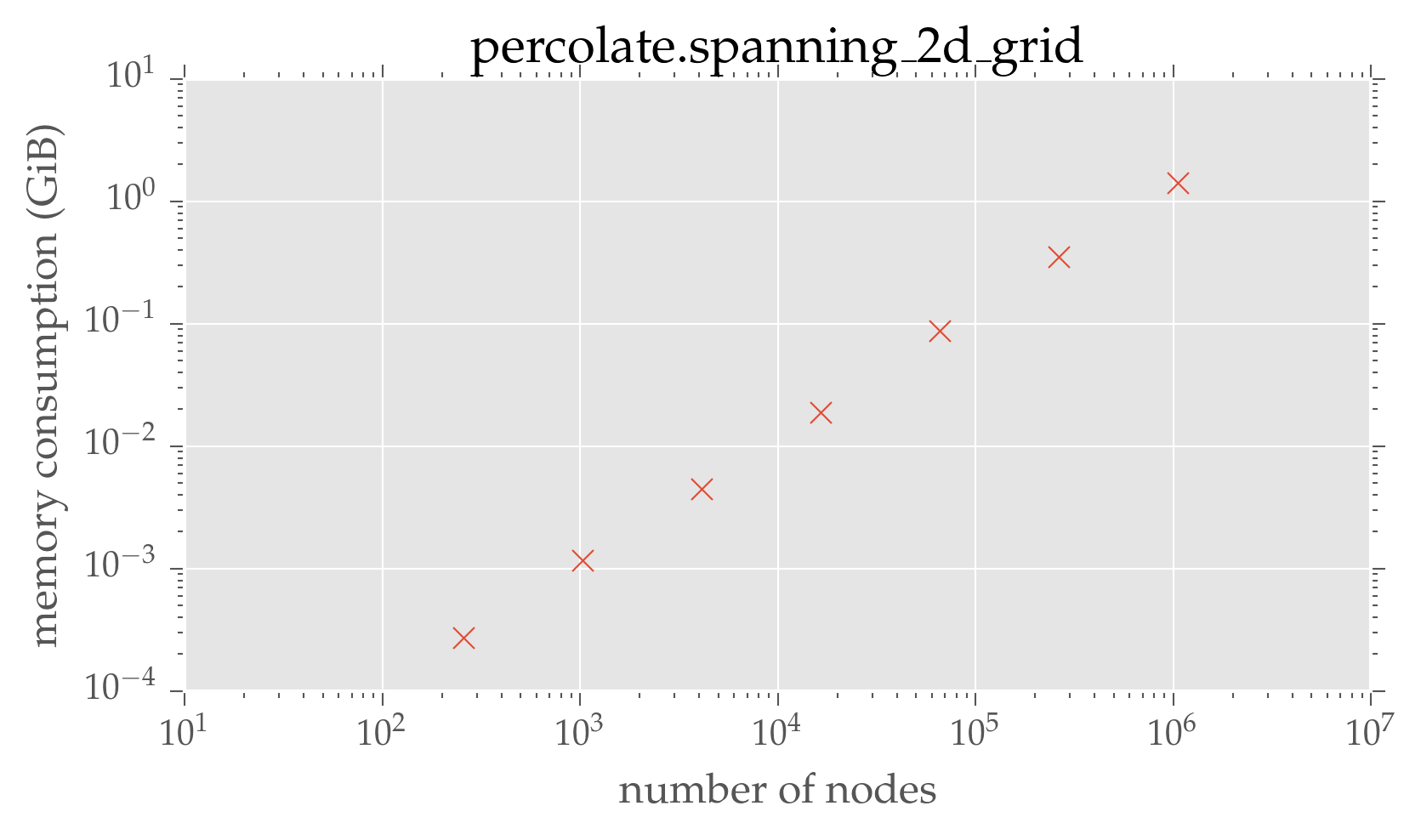
plt.loglog(dimensions ** 2, graph_mem / 2 ** 10, 'x')
plt.xlabel(r'number of nodes')
plt.ylabel('memory consumption (GiB)')
plt.title('percolate.spanning\_2d\_grid')
plt.show()
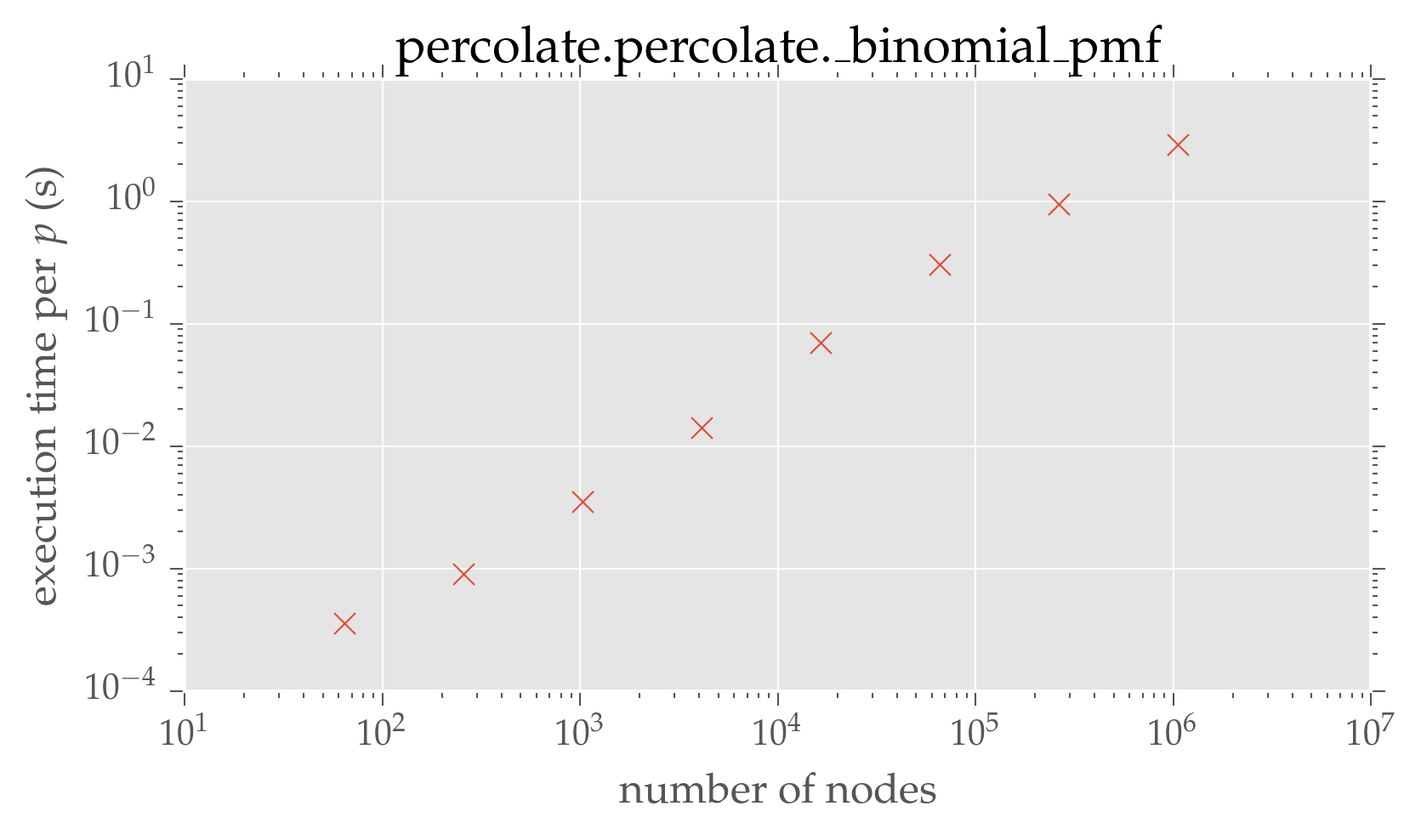
plt.loglog(dimensions ** 2, convolution_times, 'x')
plt.xlabel(r'number of nodes')
plt.ylabel(r'execution time per $p$ (s)')
plt.title('percolate.percolate.\_binomial\_pmf')
plt.show()
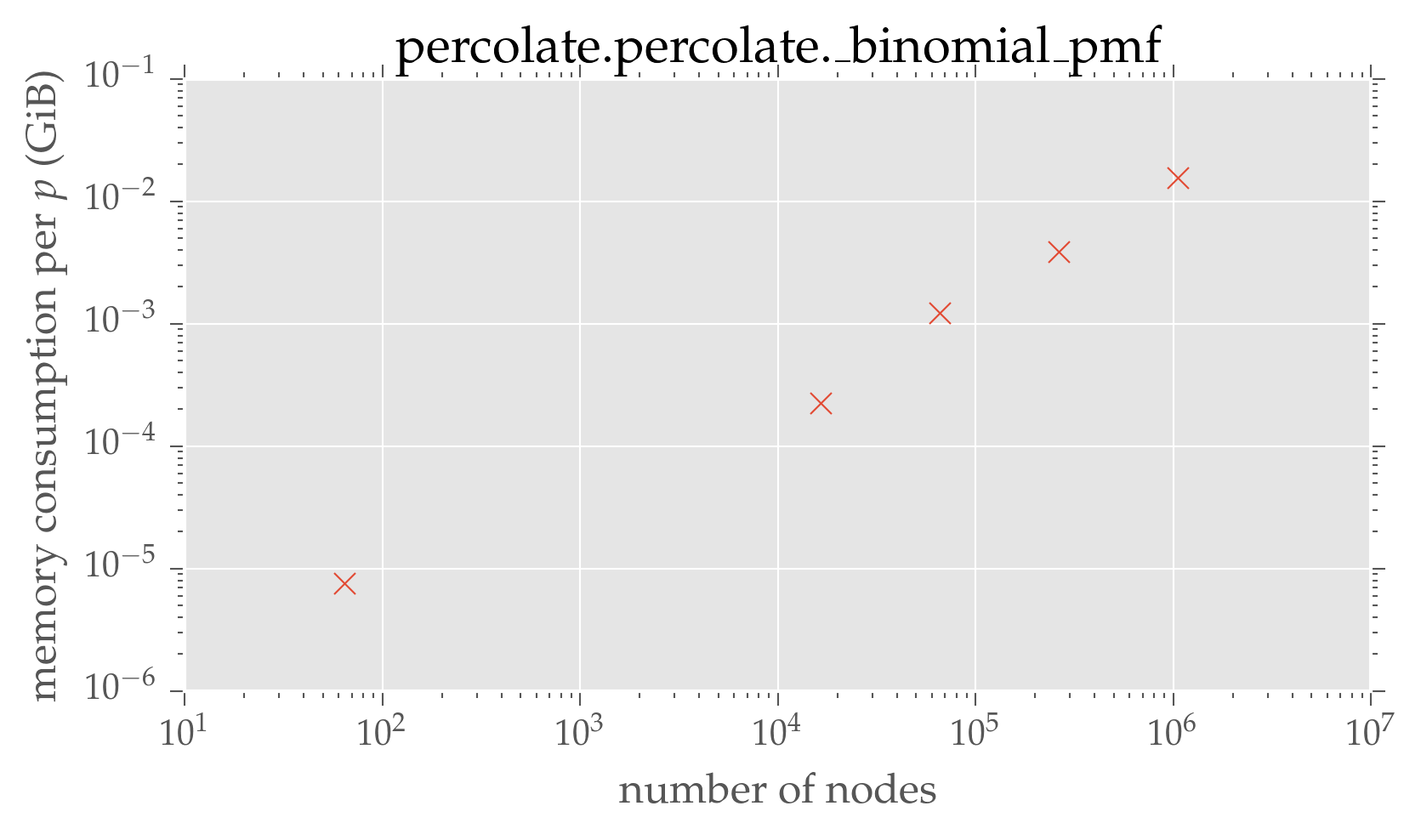
plt.loglog(dimensions ** 2, convolution_mem / 2 ** 10, 'x')
plt.xlabel(r'number of nodes')
plt.ylabel(r'memory consumption per $p$ (GiB)')
plt.title('percolate.percolate.\_binomial\_pmf')
plt.show()
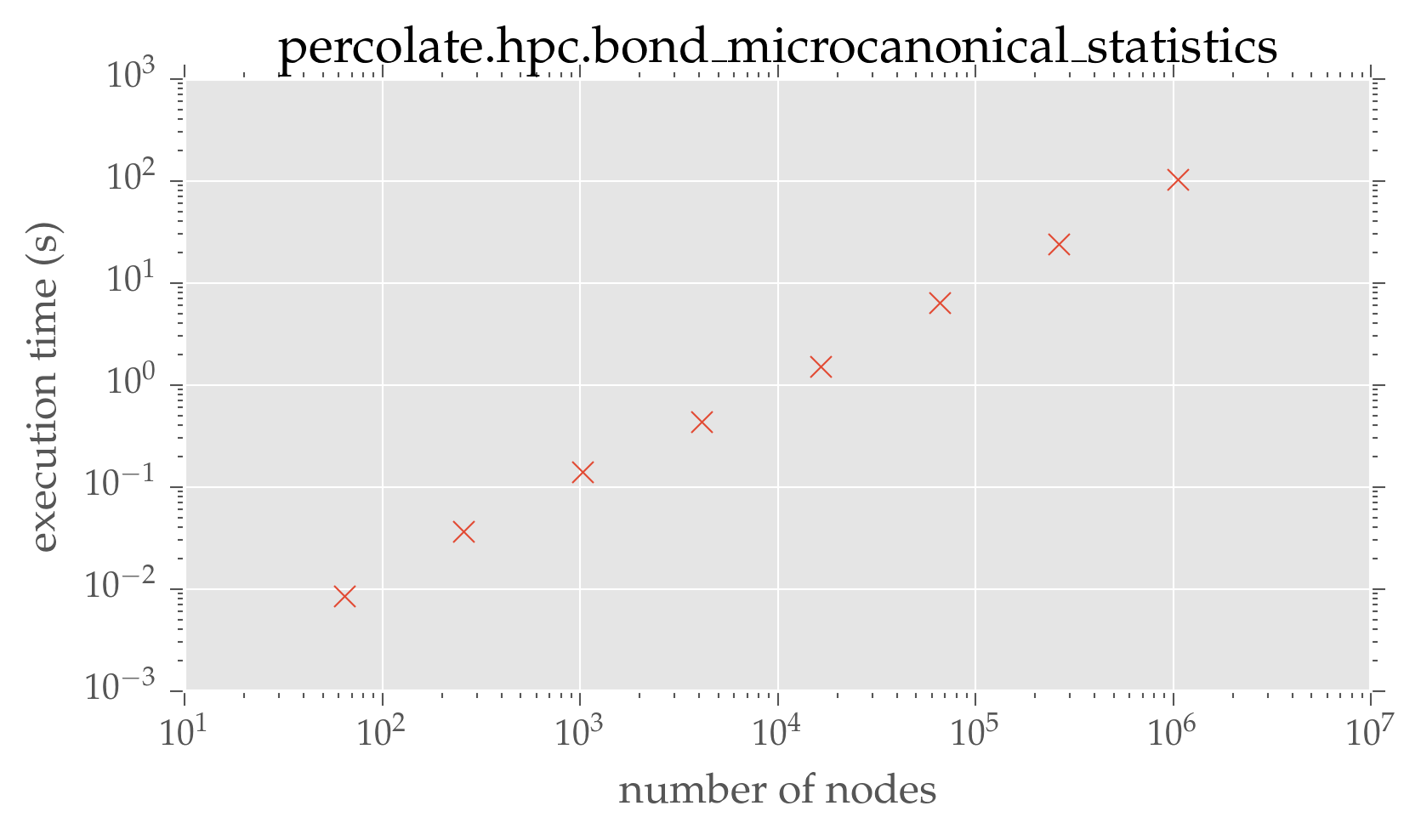
plt.loglog(dimensions ** 2, run_times, 'x')
plt.xlabel(r'number of nodes')
plt.ylabel(r'execution time (s)')
plt.title('percolate.hpc.bond\_microcanonical\_statistics')
plt.show()
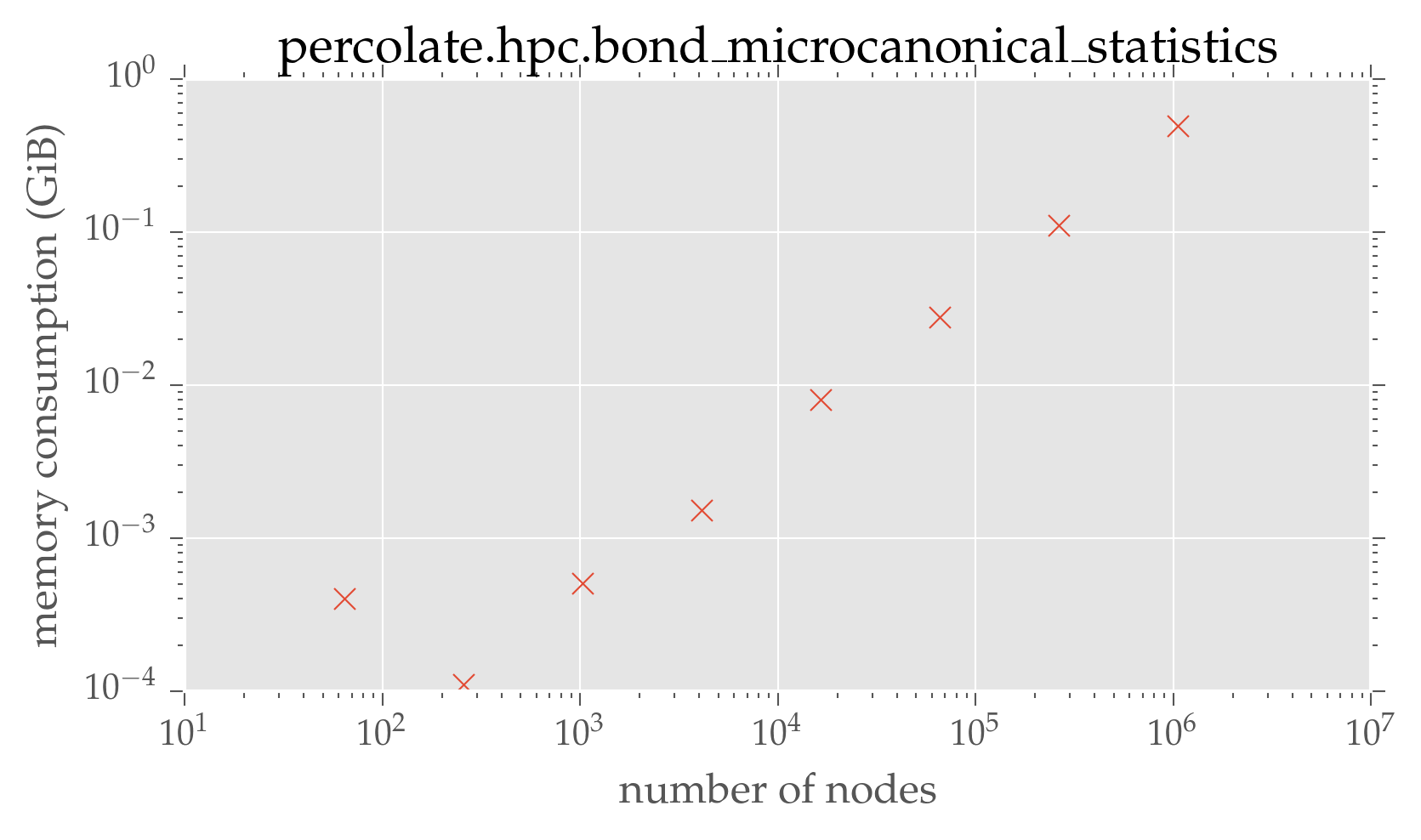
plt.loglog(dimensions ** 2, run_mem / 2 ** 10, 'x')
plt.xlabel(r'number of nodes')
plt.ylabel(r'memory consumption (GiB)')
plt.title('percolate.hpc.bond\_microcanonical\_statistics')
plt.show()